
If Java 6 is not installed, just update it or download and install it from here. 
Ensure that Java 6 is installed using this test web page.Requirements and Installation for Cytoscape 2.x 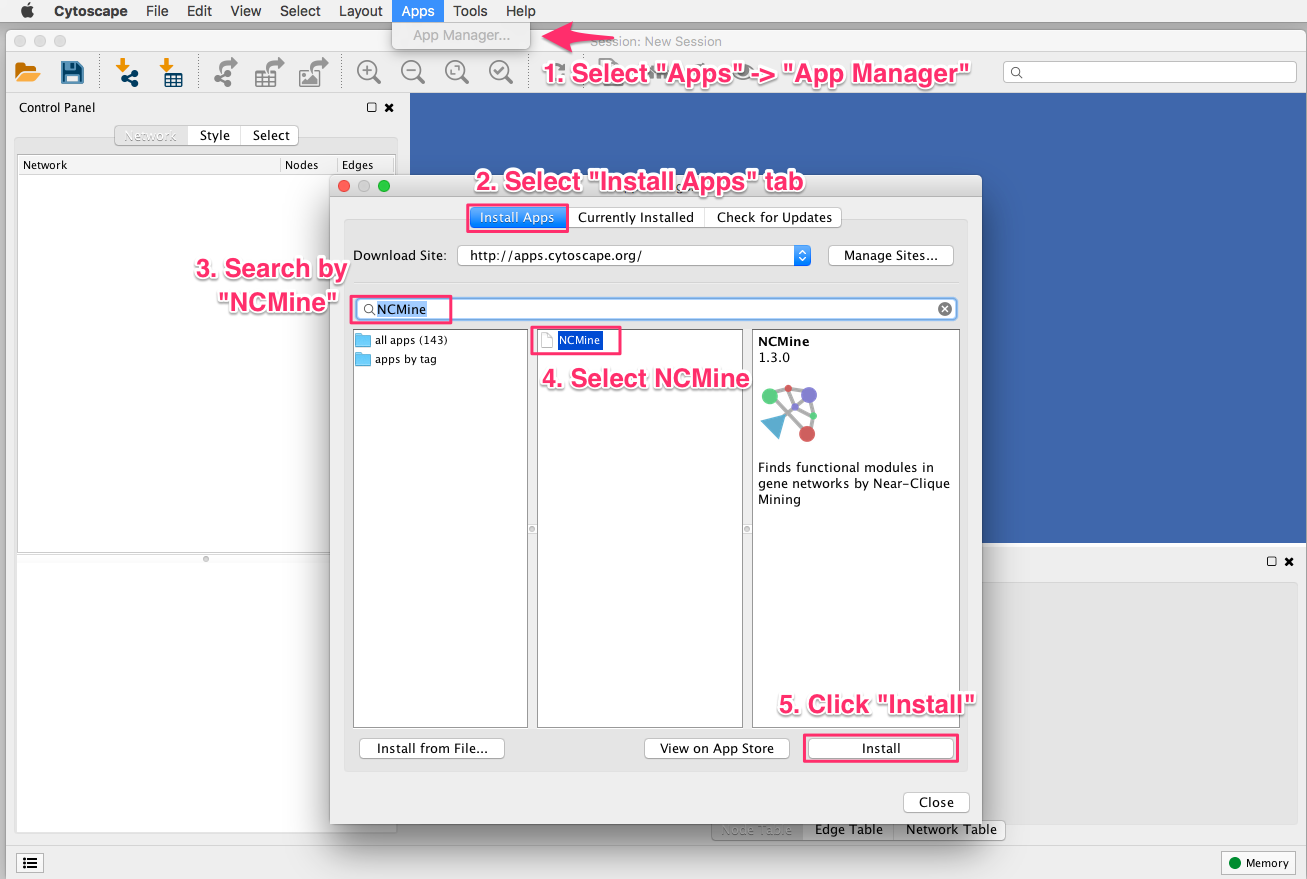
If no other path is specified in the structureViz Settings, this path is used for launching Chimera next time. Once Chimera has been started successfully, the last used path is saved in the session file.Go to Apps → structureViz → Launch Chimera.These settings are specific to each network in Cytoscape. If not using the default Chimera installation directory, set the path to the Chimera executable file in Apps → structureViz → Settings.Launch UCSF Chimera from within Cytoscape:.To check if the installation was successful, go to Apps to find the RINalyzer and structureViz menu options.Repeat the same action with the structureViz.jar file.button and select the RINalyzer.jar file. Start Cytoscape and go to the App Manager ( Apps → App Manager).Download the RINalyzer 2 app from the RINalyzer download page.Download the structureViz 2 app from the Cytoscape App Store.Install RINalyzer and structureViz app:.UCSF Chimera: Download and install the latest UCSF Chimera version (1.8 and above) from the UCSF Chimera download page.Cytoscape: Download and install the latest Cytoscape version (3.1.0 and above) from the Cytoscape download page.If Java is not installed, just update it or download and install it from here. Java: Ensure that Java 6 or 7 is installed using this test web page.CytoSEED is released under the GNU General Public License.Requirements and Installation for Cytoscape 3.x If you have any questions, problems, or suggestions, please contact dejongh. Restart Cytoscape and repeat the installation procedure above with the new version of CytoSEED. Open the Analysis folder and select "CytoSEED v.1.x" and click Delete. Type "cytoseed" in the search box and click on Search. In Cytoscape, use the "Plugins->Manage Plugins" menu item. TO UNINSTALL CYTOSEED v.1.0 IN ORDER TO INSTALL A NEW VERSION: Ctrl+Shift+T - Tile all maps (Tiled view of all networks) Ctrl+Shift+Z - Zoom out to display entire current network Ctrl+Shift+L - Load model and create session If there are problems with the download, you can restart it at any time from the Plugins -> SEED -> Download KGML Files From KEGG menu item. Next you will be prompted to download the KGML files from KEGG click on "Yes". Select the Plugins -> SEED -> Set Location of CytoSEED Folder menu item, and choose a location where the plugin can create a new folder named CytoSEED that will store the model and KGML data. Click on the arrow beside the "Network and Attribute I/O" folder, select the latest version of the KGMLReader plugin, and click on "Install".Ĥ) If you installed both plugins correctly, you will now see new menu items listed under the Cytoscape Plugins menu as KGML Reader and SEED. 
Click on the Plugins -> Manage Plugins menu item in the box labeled "Enter key words to search", type "kgml", then click on the "Search" button. Start the Cytoscape application and install the CytoSEED plugin by clicking on the Plugins -> Install Plugin from File menu item then browse to and select the CytoSEEDplugin.jar file that you downloaded.ģ) In order to format the KEGG maps properly, you must also install the KGMLReader plugin, which is available from within the Cytoscape application. The CytoSEED plugin is not compatible with earlier versions of Cytoscape.Ģ) Download the CytoSEED plugin from. SEE BELOW FOR DIRECTIONS FOR UNINSTALLING PREVIOUS VERSIONS AND INSTALLING VERSION 1.5.Ĭytoscape Model SEED Plugin (CytoSEED) READMEġ) Download and install Cytoscape 2.8.3 from. VERSION 1.5 OF THE CYTOSEED PLUGIN WAS RELEASED ON FEBRUARY 24, 2015.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
